MUSCLE: a multiple sequence alignment method with reduced time and space complexity | BMC Bioinformatics | Full Text

Jos de Bruijne a Twitter: ""A Gap in the Lower Main Sequence Revealed by #GaiaDR2" https://t.co/8pBxPkLR4o "The gap presents a diagonal feature that dips toward lower luminosities at redder colors. The gap

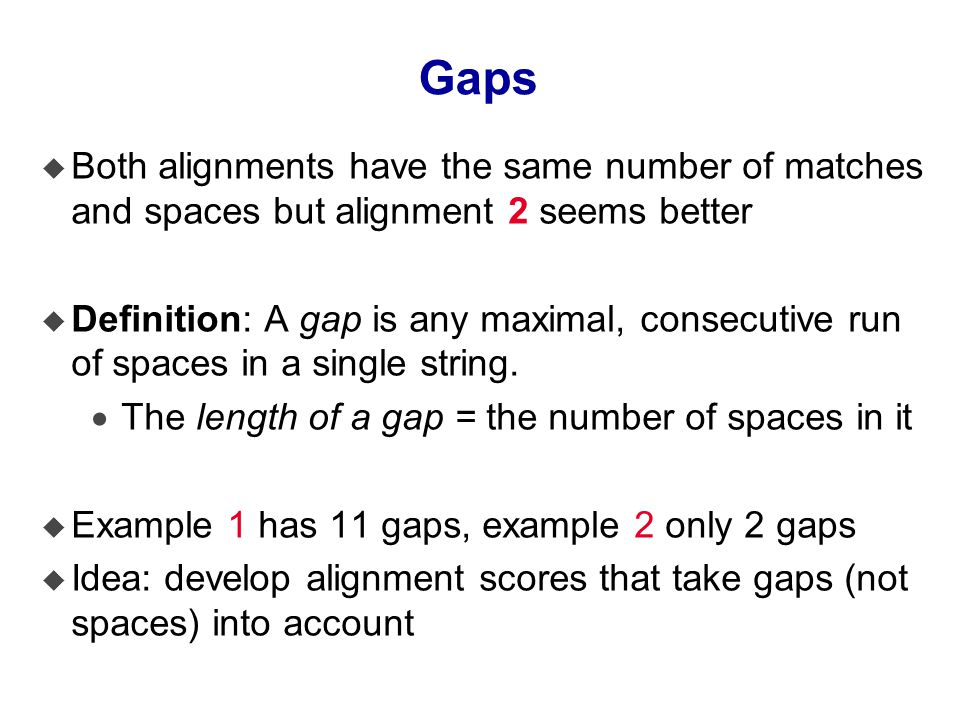
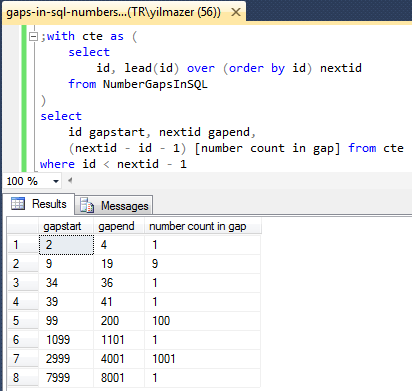
Exploring the Effects of Gap-penalties in Sequence-alignment Approach to Polymorphic Virus Detection

Protein de novo sequencing by top-down and middle-down MS/MS: Limitations imposed by mass measurement accuracy and gaps in sequence coverage - ScienceDirect

Figure 1 from Introducing Variable Gap Penalties into Three-Sequence Alignment for Protein Sequences | Semantic Scholar

Phylogeny-Aware Gap Placement Prevents Errors in Sequence Alignment and Evolutionary Analysis | Science